How pH and Salt Concentrations Shape the Self‑Assembly of Bacterial Microcompartment Shell Proteins
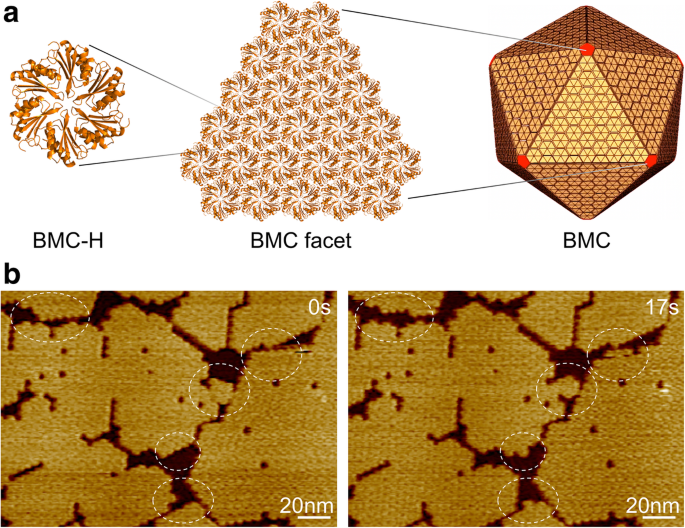
Bacterial microcompartments (BMCs) are proteinaceous organelles that sequester metabolic enzymes within a semi‑permeable shell, enabling bacteria to carry out key biochemical reactions in the absence of membrane‑bound organelles. The shell is assembled from multiple homologous proteins—hexameric BMC‑H, pentameric BMC‑P, and trimeric BMC‑T—arranged into a highly ordered icosahedral lattice. Recent efforts to engineer BMCs as nanobioreactors have highlighted the need to understand the factors that govern shell assembly. Here we use high‑speed atomic force microscopy (HS‑AFM) to quantify the dynamics of BMC‑H self‑assembly under varying pH and ionic strengths. We find that a 400‑mM salt concentration promotes the formation of large, single‑layered patches, while neutral pH (≈7) maximizes the rate of hexamer addition and removal. Moreover, we capture the transition from hexameric sheets to fiber‑like arrays, underscoring the intrinsic flexibility of these proteins. Our findings provide a detailed map of environmental conditions that stabilize BMC shell assemblies, informing rational design of synthetic BMCs and related nanomaterials. Bacterial microcompartments (BMCs) are protein‑based organelles that resemble viral capsids and are ubiquitous across bacterial phyla. They encapsulate metabolic enzymes within a semi‑permeable protein shell, thereby partitioning biochemical pathways in organisms that lack membrane‑bound organelles. The shell comprises three structural classes of proteins: BMC‑H (Pfam00936), BMC‑T (two Pfam00936 domains), and BMC‑P (Pfam03319). BMC‑H hexamers tile the facets of the shell, BMC‑P pentamers cap the vertices, and BMC‑T pseudohexamers modulate permeability. Bacterial microcompartment, shell organization, and self‑assembly. a Hundreds of BMC shell protein homologs self‑assemble into an icosahedral organelle. BMC‑H proteins, shown in yellow, form the facets; BMC‑P proteins, in red, occupy the vertices. b AFM topographs of shell facets composed of Hoch_5815 BMC‑H hexamers. Dynamic events (circles) were observed within seconds using HS‑AFM. Specific protein‑protein interactions dictate the precise architecture of the BMC shell. Lateral contacts between hexamers are the primary determinants of shell geometry. BMC‑H homologs can assemble into two‑dimensional sheets, nanotubes, or filamentous structures, illustrating the versatility of this protein family. Because of their modularity, BMCs have emerged as promising scaffolds for nanobioreactors, metabolic engineering, and drug delivery. Despite extensive structural characterization via X‑ray crystallography, electron microscopy, and dynamic light scattering, the real‑time assembly dynamics of BMC shells remain poorly understood. High‑speed AFM has recently enabled visualization of BMC‑H self‑assembly in solution, revealing both assembly and disassembly events at sub‑second resolution. In this study, we extend these observations by systematically probing how pH and salt concentration influence the size, stability, and dynamics of BMC‑H patches. The purified BMC‑H protein (Hoch_5815) from Haliangium ocraceum was kindly provided by Dr. Kerfeld (Lawrence Berkeley National Laboratory). Stock samples (~80 mg mL−1 in 50 mM Tris‑HCl, pH 7.8, 100 mM KCl, 10 mM MgCl2) were diluted to 0.5 mg mL−1 in the desired buffer before imaging. The control buffer consisted of 50 mM Tris‑HCl (pH 7.8) and 10 mM MgCl2. Samples were adsorbed onto freshly cleaved mica for 5 min, rinsed with the corresponding buffer, and imaged in solution using a JPK NanoWizard ULTRA speed AFM. Scanning was performed at 30 or 40 Hz in AC mode with an ULTRA Speed 2.8 µm scanner and Ultra‑Short Cantilever USC‑0.3 MHz probes. A minimal loading force (~100 pN) was applied to preserve native protein interactions. Initial image analysis used JPK SPM Data Processing. For dynamic studies, a custom macro in Image SXM was employed. Patch sizes were extracted from 512 × 512 pixel images at 30 Hz: images were flattened, thresholded, and binarized to distinguish protein from background. Particle analysis yielded the surface area of individual patches. Adjacent patches were separated by >3 pixels (~2 nm). Dynamic events were quantified from 256 × 256 pixel series at 40 Hz (≈6.4 s per frame). Binary difference images highlighted added or removed hexamers; the number of events (N) was divided by the total protein area (A) and elapsed time (T) to obtain the dynamic rate R: R = N / (A × T) Data are presented as mean ± SD. Statistical significance was assessed via multivariate or two‑way ANOVA as appropriate. Hoch_5815 hexamers, expressed in E. coli and characterized by six‑fold symmetry, assemble into single‑layered sheets within seconds— the fundamental building blocks of the icosahedral BMC shell. HS‑AFM reveals both the spatial organization of these sheets and the kinetics of their growth and dissolution. Patch size increased progressively from pH 3 to 10, indicating that alkaline conditions favor hexamer association. At neutral pH 7, the dynamic rate peaked, while rates dropped sharply in both acidic and alkaline environments. These trends differ from the behavior of RmmH, which forms nanotubes at pH 8 but disassembles at pH 10. Effects of environmental pH on the self‑assembly of Hoch_5815. a Mean surface areas of individual patches (n = 50). b Mean dynamic event rates (n = 50). *p < 0.05, ns not significant (multivariate ANOVA). Electrostatic interactions at the hexamer–hexamer interface likely underlie the pH dependence. Acidic conditions reduce hexamer association, whereas alkaline conditions stabilize the sheets but reduce the overall dynamic exchange. Neutral pH provides an optimal balance between stability and flexibility. Increasing ionic strength from 100 to 500 mM (CaCl2, MgCl2, KCl) led to progressively larger patches. At 500 mM, double‑ or multilayered sheets were observed. A sharp increase in patch size occurred between 200 and 300 mM, especially for CaCl2, suggesting a heightened sensitivity to calcium. Dynamic rates decreased with higher salt, reflecting stronger lateral interactions and reduced hexamer turnover. Effects of salt concentration on the self‑assembly of Hoch_5815. a Mean patch areas under 100–500 mM CaCl2, MgCl2, KCl (n = 50). Changes were significant between 200 and 300 mM (***p < 0.001, *p < 0.05, ns not significant, two‑way ANOVA). b Mean dynamic event rates under the same conditions. Each 100 mM increment produced a significant change in dynamics (***p < 0.001, *p < 0.05, ns not significant, two‑way ANOVA). Notably, 400 mM of any of the cations produced the largest, most stable single‑layer assemblies with the highest proportion of assembly over disassembly events. At 500 mM, the reduced dynamics and the presence of multilayer sheets indicate highly stable interactions. Using a reduced scan force (~100 pN), we captured both assembly and disassembly events at the single‑hexamer level. Under pH 7.5 with only 10 mM MgCl2, we observed occasional fiber‑like structures co‑existing with sheet disassembly. These fibers, with a spacing of 3.72 ± 0.31 nm and a height of 2.46 ± 0.22 nm, are shorter than the 3.45 ± 0.16 nm height of intact hexamers, suggesting they arise from individual monomers or partially disassembled oligomers. Formation and dynamics of fibrous structures alongside shell sheet assemblies. a Fiber appearance during sheet disassembly. b AFM topograph of fibers. c Cross‑section shows 3.72 ± 0.31 nm spacing (n = 30). d Topograph of hexameric patches. e Cross‑section reveals 3.45 ± 0.16 nm height (n = 25). f Fibers are significantly shorter than sheets (*p < 0.05, two‑way ANOVA). g Proposed model: fibers consist of monomeric Hoch_5815 chains. Our quantitative HS‑AFM analysis demonstrates that BMC‑H self‑assembly is exquisitely sensitive to environmental cues. Neutral pH and moderate ionic strength (≈400 mM) create conditions that maximize both the size and dynamic turnover of hexameric patches, mirroring the cytosolic milieu of many bacteria (pH 7.4–7.8; ion concentration 100–400 mM). This aligns with prior observations that shell proteins can assemble across a broad pH range but exhibit optimal dynamics at physiological conditions. Comparisons with other BMC types, such as carboxysome shell proteins (CcmK), reveal common themes: increased salt promotes larger, more ordered assemblies, while excessive ionic strength can lead to multilayer formation or disassembly of tubular structures. The distinct responses of Hoch_5815 to CaCl2 versus MgCl2 suggest that divalent cations modulate inter‑hexamer contacts through electrostatic screening or specific binding. These insights are directly applicable to the rational design of engineered BMCs. By tuning pH and ionic strength during assembly, one can bias the shell toward desired architectures—single‑layer sheets for maximal enzyme encapsulation or multilayered shells for structural reinforcement. Moreover, the observed fiber‑like intermediates may serve as templates for novel nanofibers or could be harnessed to modulate shell permeability. Future work should integrate computational modeling of hexamer–hexamer interfaces with experimental AFM data to dissect the thermodynamic landscape of BMC assembly. Such combined approaches will deepen our understanding of how natural BMCs adapt to fluctuating cellular environments and will guide the construction of robust, programmable nanostructures. High‑speed AFM has allowed us to map the self‑assembly landscape of BMC‑H hexamers under physiologically relevant conditions. Large, stable single‑layer patches are favored at 400 mM salt, while neutral pH enhances the dynamic exchange of hexamers. The discovery of fiber‑like assemblies further highlights the inherent plasticity of BMC shell proteins. These findings lay a quantitative foundation for engineering BMC‑based nanobioreactors and other biomaterials that can be tuned by simple changes in pH or ionic strength. Bacterial microcompartment Bacterial microcompartment hexamer Bacterial microcompartment pentamer Bacterial microcompartment trimer Dynamic light scattering Escherichia coli Electron microscopy High-speed atomic force microscopyAbstract
Introduction
Methods
Sample Preparation
Atomic Force Microscopy
Image Processing and Analysis
Results
Response to pH Variation
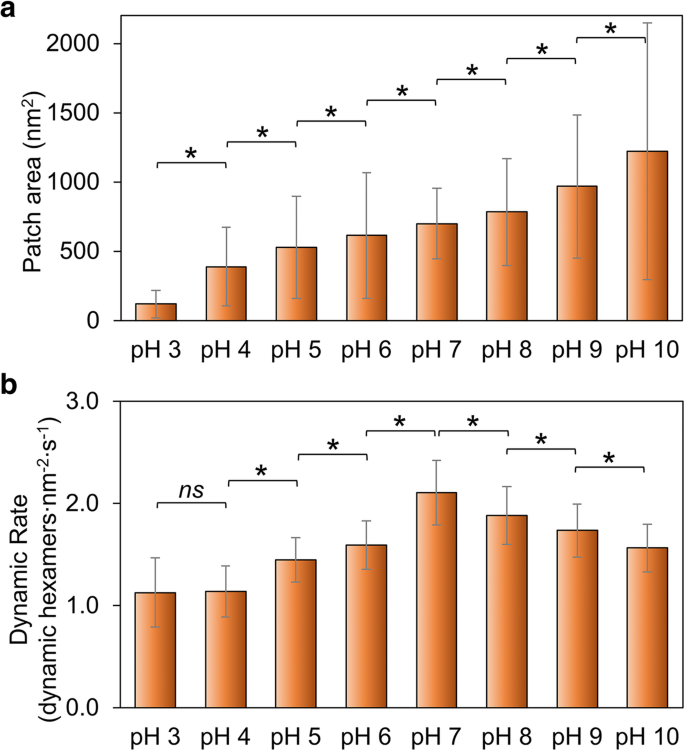
Response to the Variation of Salt Concentrations
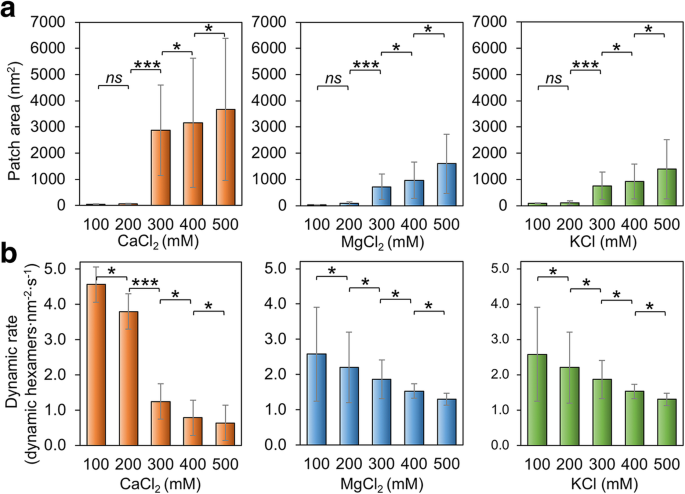
Flexibility of BMC‑H Protein Assembly
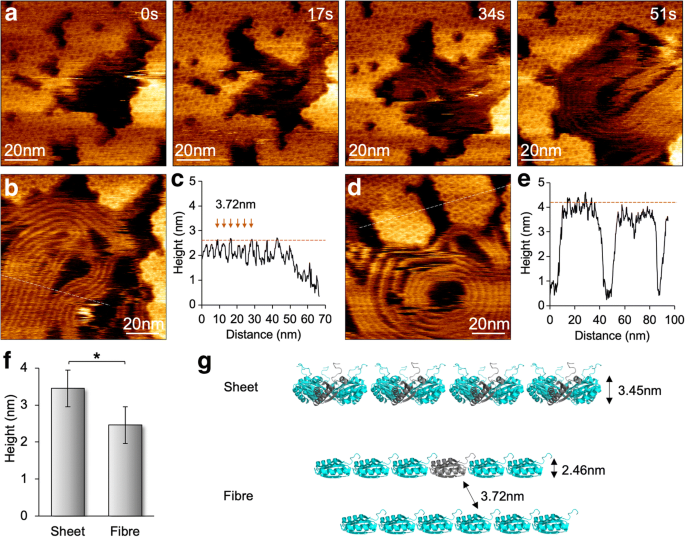
Discussion
Conclusions
Abbreviations
Nanomaterials
- Secure, On‑Demand Private LTE: IPX Network Innovation & Dynamic Bandwidth
- First‑Principles Investigation of the Structural Stability and STM Imaging of Borophene on Ag(111)
- Easily Synthesized Wormhole‑Like Mesoporous SnO₂ via Evaporation‑Induced Self‑Assembly: Superior Ethanol Gas‑Sensor Performance
- First-Principles Analysis of Point Defects in GaAs/AlAs Superlattices: Stability, Band Structure, and Mobility
- Carbon Nanomaterials for Treating Heavy‑Metal‑Contaminated Water and Advancing Environmental Remediation
- Investigating Meridian Response Currents to Electrical Pulse Acupuncture: A Physics-Based Study
- Investigating ROS and Cell Cycle Arrest in the Genotoxic Effects of Gold Nanorod Core/Silver Shell Nanostructures
- How the Pandemic is Accelerating Automation: Robots, Cobots, and Smart Manufacturing
- Manufacturing Recovery: Transforming Talent Acquisition & Retention for the Future
- Automation and Robotics: Redefining the Future of Industry



