Magnetic Graphene Field‑Effect Transistor Biosensor for Ultra‑Sensitive Single‑Strand DNA Detection
In this work, we present a magnetic graphene field‑effect transistor (MGFET) biosensor fabricated by transferring a chemical‑vapor‑deposition (CVD) graphene film onto a glass substrate to form the conductive channel and sensing layer. The graphene surface is functionalized with 1‑pyrenebutanoic acid succinimidyl ester (PBASE), which acts as a molecular anchor for immobilizing a probe aptamer. Magnetically labeled complementary single‑strand DNA (cDNA) is captured via hybridization with the aptamer, and the interaction is transduced into a measurable impedance change. Under a periodically modulated magnetic field, the MGFET impedance exhibits oscillations whose amplitude is directly proportional to the cDNA concentration. Using this principle, the sensor detects single‑strand DNA with a detection limit of 1 pM. The observed behavior is rationalized by a model in which the magnetic force bends the DNA‑bead complex, thereby modulating the double‑layer capacitance of the graphene channel. By applying a periodic magnetic field, the impedance can be sampled repeatedly, allowing signal‑to‑noise ratio (SNR) enhancement through integration over multiple periods. This study demonstrates a highly sensitive DNA biosensor and suggests its potential applicability to other biomolecules and cellular assays. The detection of DNA is pivotal for molecular biology research and the diagnosis of genetic disorders [1,2,3]. Over the past decade, biosensors based on fluorescence [4,5], electrochemistry [6–9], and field‑effect transistors (FET) [10–13] have been developed for this purpose, with FET‑type sensors gaining particular attention due to their inherent sensitivity and specificity. Kaisti et al. introduced a peptide‑nucleic‑acid‑functionalized FET for unlabeled single‑strand DNA detection [12], while Kim et al. fabricated a complementary‑metal‑oxide‑semiconductor (CMOS)‑based DNA charge sensor [13].
Graphene, with its exceptional surface area, electrical conductivity, and carrier mobility, has emerged as an ideal platform for constructing FET biosensors [14–16]. Cai and colleagues reported a graphene FET (GFET) capable of ultrasensitive DNA detection via peptide‑nucleic‑acid hybridization [15], and our group previously demonstrated a multi‑channel GFET for studying DNA binding kinetics and base‑pair mismatches [16].
In conventional GFETs, an external gate electrode generates an electric field that establishes a double‑layer at the graphene–electrolyte interface [17–19], thereby modulating device conductance. Recent studies have achieved femtomolar detection limits with GFETs [20,21]; however, these high sensitivities rely on expensive semiconductor analyzers and Ag/AgCl reference electrodes, limiting practical deployment.
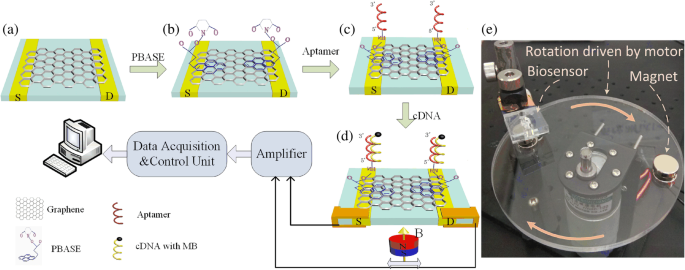
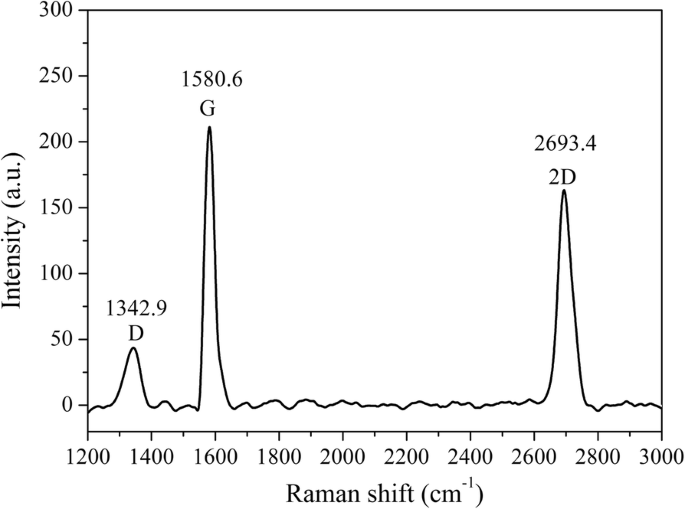
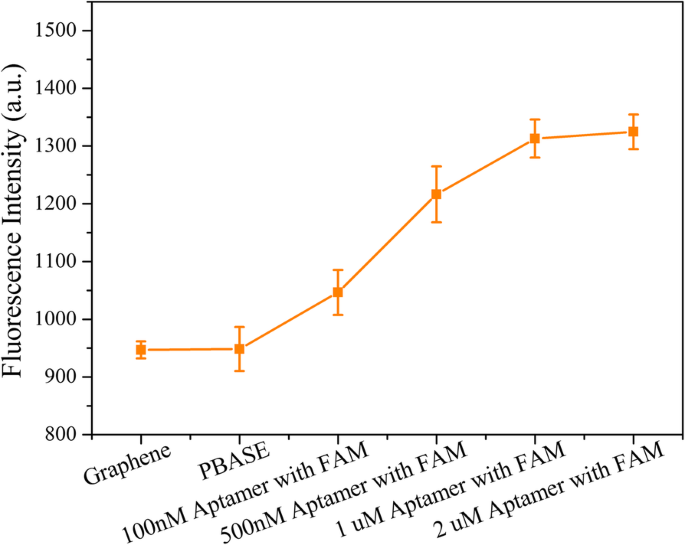
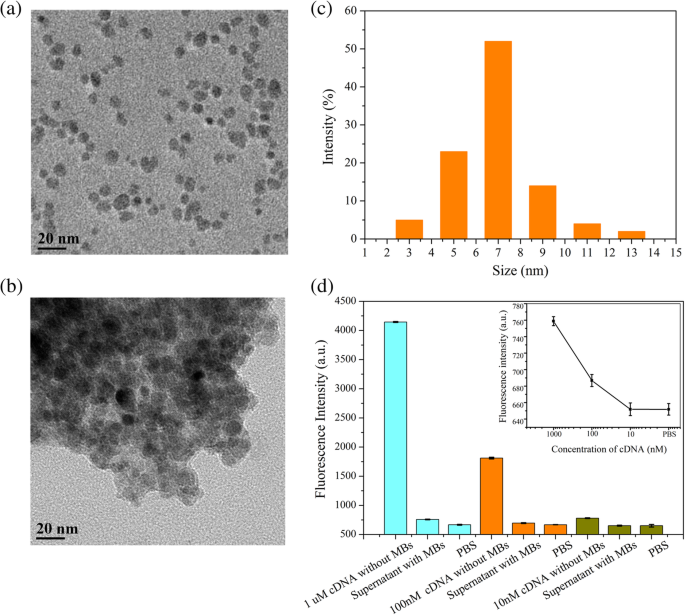
To overcome these challenges, we devised a magnetic graphene FET (MGFET) that uses a magnetic field, rather than an electric gate, to modulate the graphene channel. The MGFET incorporates a CVD graphene film transferred onto a glass substrate flanked by indium‑tin‑oxide (ITO) electrodes. The graphene is functionalized with PBASE, enabling covalent attachment of a probe aptamer that captures magnetically labeled cDNA. When subjected to a periodic magnetic field, the MGFET impedance oscillates in a manner that correlates with cDNA concentration. This magnetic approach eliminates direct electrode contact, simplifying integration and broadening applicability. The following sections detail the fabrication, detection system, and underlying detection mechanism of this novel MGFET biosensor. A glass substrate with ITO electrodes was sourced from Hua Nan Xiang Cheng Ltd. (China). The probe aptamer, cDNA, and mismatched DNA were purchased from Sangon Biotech Inc. (Shanghai, China). The aptamer sequence is (5′‑NH2‑TGG ACC CCC TCA TAA CGC CTC CTT TTC‑FAM‑3′); the complementary DNA (cDNA) sequence is (5′‑NH2‑GAA AAG GAG GCG TTA TGA GGG GGT CCA‑3′); the fully mismatched DNA is (5′‑NH2‑TCC CCT TCT TAT GGC CTG TTT TTC AAC‑3′); and the single‑base mismatched DNA is (5′‑NH2‑GAA AAG GAG TCG TTA TGA GGG GGT CCA‑3′). PBASE and dimethyl sulfoxide (DMSO) were obtained from Sigma‑Aldrich (Shanghai, China). Magnetic nanobeads (MBs) with carboxyl functionalization (10 mg mL−1) were acquired from Xianfeng Nano Material Technology Co., Ltd. (Nanjing, China). 1‑Ethyl‑3-(3‑dimethylaminopropyl) carbodiimide hydrochloride, N‑hydroxysuccinimide, sodium dodecylbenzenesulfonate (SDS), and phosphate‑buffered saline (PBS, pH 7.4) were purchased from Sigma‑Aldrich (Shanghai, China). A Raman spectrometer (SPEX‑1403, SPEX) characterized graphene quality and confirmed PBASE functionalization. Fluorescence photometry (LS55, PerkinElmer) quantified the coupling of magnetic nanoparticles to cDNA. A custom data‑acquisition system recorded real‑time MGFET impedance. After sonication for 20 min, 20 µL of 4 mg mL−1 MB suspension was mixed with 200 µL of 2 mg mL−1 EDC and 200 µL of 2 mg mL−1 NHS for 15 min to activate the MBs [22,23]. Subsequently, 20 µL of cDNA was added and incubated for 2 h at room temperature with gentle agitation. A magnetic field then enriched the MB/cDNA conjugates, which were washed three times with PBS and resuspended for downstream use. The MGFETs were fabricated in a three‑step process: (1) a CVD graphene film was transferred onto a glass plate serving as the conductive channel between the two ITO electrodes (Fig. 1a). (2) A 10 mM PBASE solution in DMSO was incubated on the graphene for 12 h at room temperature, allowing π–π stacking with the graphene lattice (Fig. 1b). Excess PBASE was removed by rinsing with DMSO and PBS. (3) A 2 µM probe aptamer solution was introduced and incubated for 4 h at room temperature, enabling covalent coupling via the NHS ester of PBASE (Fig. 1c). Finally, unbound aptamer was washed away with 0.2 % SDS three times. Functionalization and detection principle of the MGFETs. a Graphene film grown by chemical vapor deposition. b Functionalization of graphene by PBASE. c Immobilization of probe aptamer via PBASE. d Hybridization of the probe aptamer with cDNA. e Photograph of the detection device The CVD graphene film was successfully transferred onto the glass substrate, as confirmed by the Raman spectrum (Fig. 2). The presence of the G, D, and 2D peaks indicates high‑quality graphene; the I2D/IG ratio confirms a multilayer film, while the low ID/IG ratio evidences minimal defects [24–26]. Raman spectrum of the transferred graphene film Direct attachment of aptamers to pristine graphene is challenging due to the lack of functional groups; therefore, PBASE serves as a π–π anchored linker. The succinimide group of PBASE reacts with the 5′‑NH2 of the aptamer via NHS chemistry (Fig. 1c). Successful aptamer immobilization was confirmed by the fluorescence intensity of the 3′‑end FAM label (Fig. 3). Fluorescence increased with aptamer concentration until plateauing at ~2 µM, indicating surface saturation. Consequently, subsequent experiments employed a 2 µM aptamer concentration. Fluorescence characterization of aptamer immobilization. Error bar represents the standard deviation of 5 independent analyses. Transmission electron microscopy revealed that the MBs are ~7 nm in diameter (Fig. 4a,b). To maximize capture efficiency, a 4 mg mL−1 MB concentration was employed for activation. Coupling efficiency was monitored by measuring the decrease in FAM fluorescence in the supernatant (Fig. 4d). Complete enrichment of 10 nM cDNA was achieved, as evidenced by the fluorescence level of the supernatant returning to that of PBS. Characterization of MB/cDNA coupling. a TEM of MBs. b TEM of MB/cDNA conjugates. c Particle size distribution of MBs. d Fluorescence-based coupling efficiency. Error bar represents the standard deviation of 5 independent analyses. After a 10‑min incubation of MB/cDNA with the MGFETs, unbound MBs were removed by PBS washes, leaving only MB/cDNA on the sensor (Fig. 1d). A permanent magnet mounted on a rotating motor applied a periodic magnetic field (Fig. 1e). The impedance of the MGFET was recorded using a custom detection device. The relationship between magnetic field strength and MGFET impedance was investigated to identify the optimal field parameters (Fig. 5). In MGFETs, magnetic force bends the MB/cDNA complex, altering the distance between the complex and the graphene surface, which in turn modulates the double‑layer capacitance. Three regimes were observed: (1) <100 mT, where the magnetic force is insufficient to bend the DNA, yielding negligible impedance change; (2) 100–200 mT, where bending occurs rapidly, producing a pronounced impedance response; and (3) >220 mT, where the DNA rod reaches its bending limit, and impedance stabilizes (Fig. 5b). This elastic‑rod model aligns with prior reports on DNA mechanics [29,30]. Influence of magnetic field intensity on impedance. a Time‑domain impedance under varying field strengths. b Impedance versus magnetic field intensity. Error bar represents the standard deviation of 5 independent analyses. With a fixed magnetic field of 240 mT, impedance changes were recorded across a range of MB/cDNA concentrations (Fig. 6a). The application of the periodic magnetic field induced a clear impedance oscillation. To suppress noise, a sample‑integration algorithm (SIA) was employed: (1) data points collected during the non‑magnetic period (T0) were normalized to zero; (2) points during each magnetic period (TM) were sampled and averaged; after four cycles, the periodic impedance change was extracted (Fig. 6b). This approach effectively enhances the SNR by leveraging repeated sampling. a Time‑domain impedance fluctuations at various cDNA concentrations. b Impedance change versus cDNA concentration. The impedance change exhibited a linear relationship with cDNA concentration (Fig. 7). The high sensitivity stems from (1) the mechanical bending of the MB/cDNA complex, which amplifies modulation of the graphene double layer, and (2) the periodic magnetic field that permits SIA‑based noise suppression. By increasing the number of magnetic periods, the system’s SNR can be further improved. Relationship between MGFET impedance and target DNA concentration. Error bar represents the standard deviation of 5 independent analyses. Selectivity was evaluated using a fully mismatched DNA and a single‑base mismatched DNA, each coupled to MBs. After 10 min incubation with the MGFETs and PBS washing, the fully mismatched MB/DNA complex was largely removed, resulting in negligible impedance change (Fig. 8a), confirming high selectivity. The single‑base mismatched DNA produced an impedance change slightly lower than the fully complementary strand but higher than the non‑complementary control, demonstrating that the sensor can discriminate single‑base mismatches (Fig. 8b). While the aptamer’s intrinsic specificity governs the selectivity, the MGFET architecture contributes to the overall sensitivity. Relationship between MGFET impedance and concentration of mismatched DNA. Error bar represents the standard deviation of 5 independent analyses. We have developed a magnetic graphene FET biosensor that leverages magnetically labeled DNA and a periodic magnetic field to transduce binding events into impedance oscillations. The magnetic force mechanically bends the MB/cDNA complex, modulating the graphene double‑layer and yielding a 1 pM detection limit for single‑strand DNA. The periodic magnetic field enables signal‑integration based noise suppression, further enhancing sensitivity. Beyond DNA detection, this platform offers a versatile framework for measuring the mechanical properties of nucleic acids and for detecting other biomolecules or cells. All data generated or analyzed during this study are included within the article. Complementary magnetically labeled single‑strand DNA Chemical vapor deposition Dimethyl sulfoxide Field‑effect transistor Graphene field‑effect transistor Magnetic nanobeads Magnetic graphene field‑effect transistor N‑Hydroxysuccinimide 1‑Pyrenebutanoic acid succinimidyl ester Sodium dodecyl sulfate phosphate‑buffered saline Sodium dodecylbenzenesulfonate Sample integration algorithm Transmission electron microscopy Indium tin oxideAbstract
Introduction
Methods
Materials and Instrument
Coupling cDNA to MBs
Fabrication of MGFETs
Results and Discussion
Characterization of MGFETs
Characterization of MB/cDNA
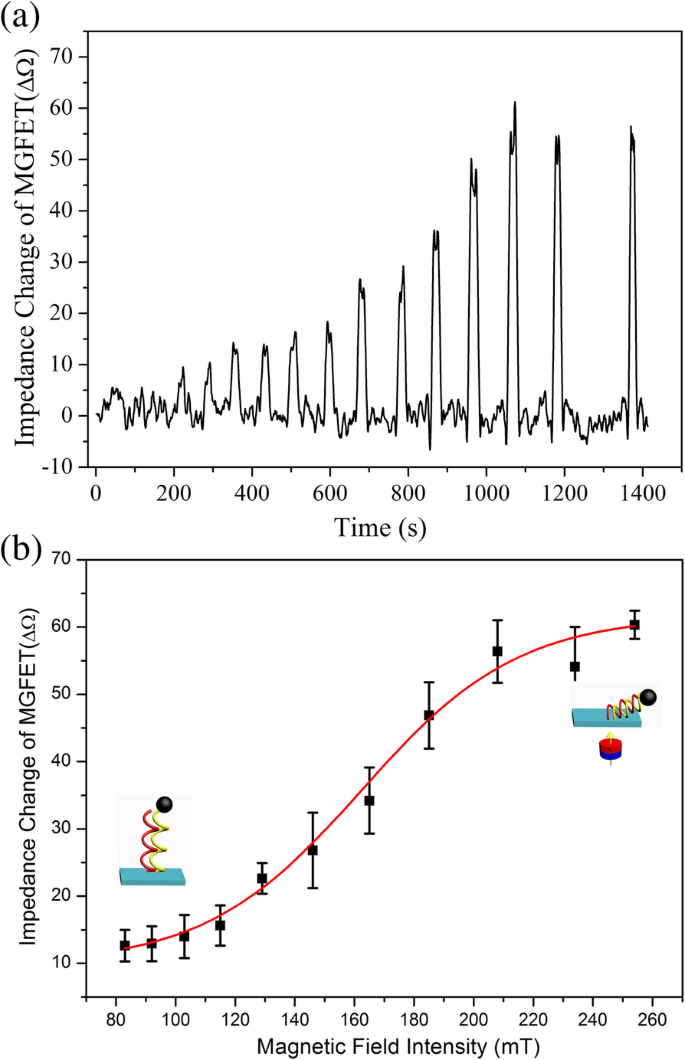
Analysis of Magnetic Field Intensity
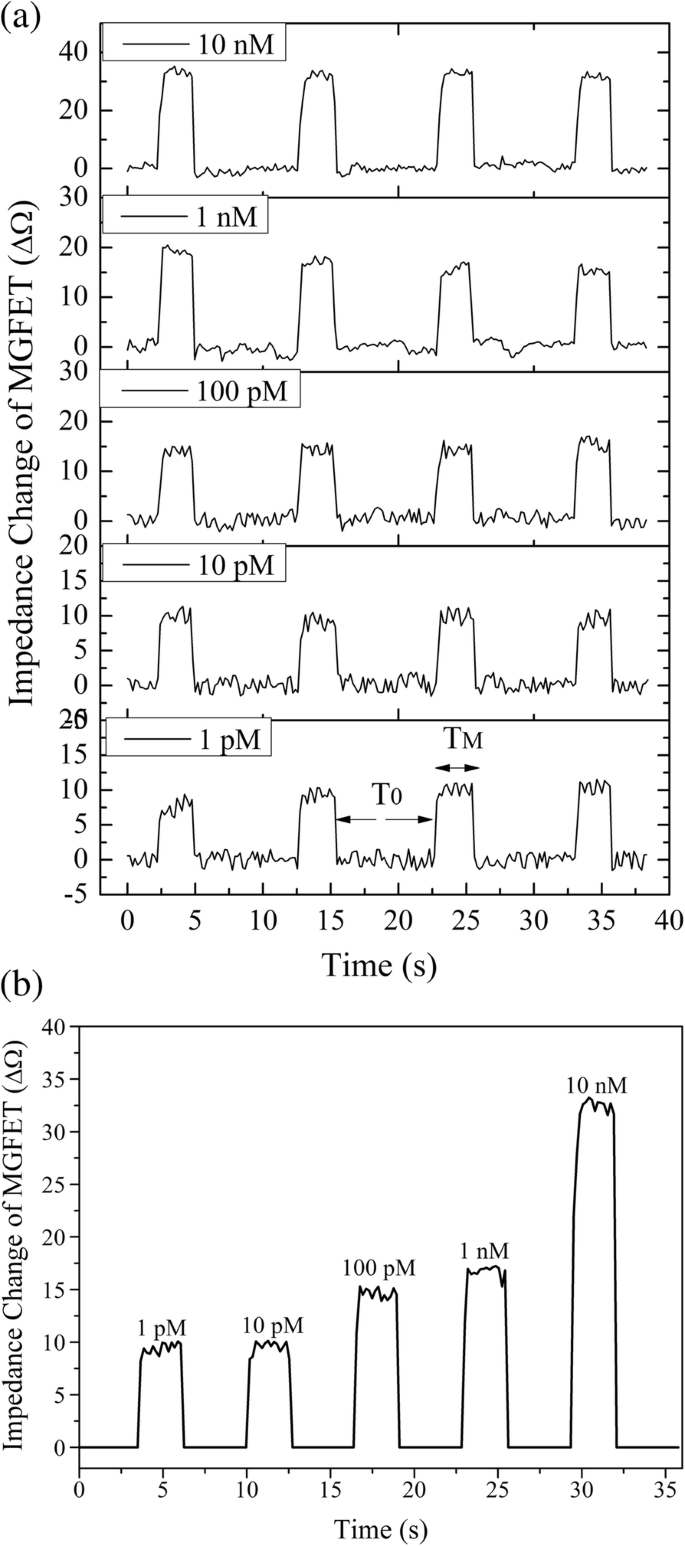
Detection of cDNA
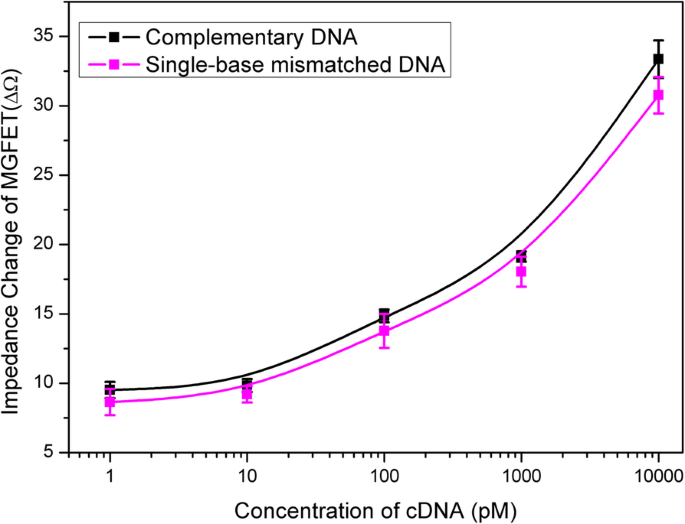
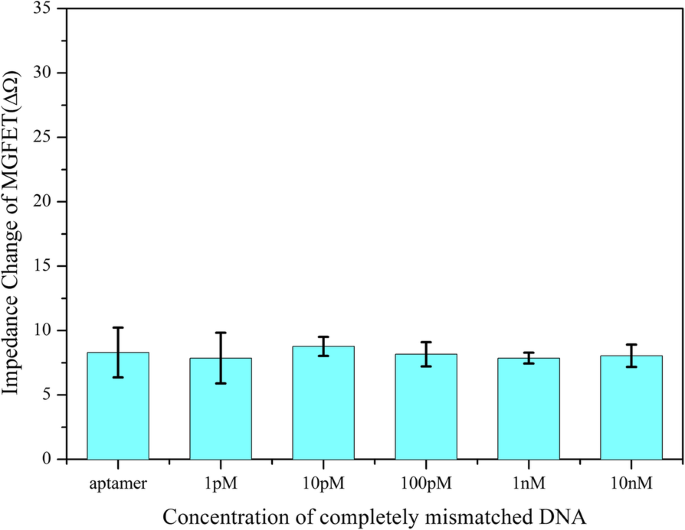
Selectivity of the MGFETs
Conclusions
Availability of Data and Materials
Abbreviations
Nanomaterials
- Nanodiamonds: Ultra‑Precise Magnetic Sensors for Biomedical and Quantum Applications
- A Flexible Graphene Biosensor for Rapid, Highly Sensitive Detection of Ovarian Cancer Cells
- Ultra‑Sensitive Electrochemical DNA Biosensor for Vibrio cholerae Detection Using Polystyrene‑Acrylic Acid Nanoparticles and Gold Nanoparticles
- Hybrid Graphene/WO₃ and Graphene/CeOx Electrodes for High‑Performance Supercapacitors
- High-Sensitivity Graphene Oxide Fluorescent Aptasensor for Rapid Detection of CCRF-CEM Leukemia Cells
- Large‑Area, Uniform Graphene Nanomeshes Fabricated for High‑Speed, Room‑Temperature Terahertz Detection
- Enhanced Formaldehyde Detection at ppb Levels Using Reduced Graphene Oxide–Coated Silicon Nanowire Sensors
- Dual‑Target Magnetic Nanoparticles Enable High‑Purity Lymphatic Endothelial Cell Isolation and Dual‑Modality Imaging in Colorectal Cancer
- Advanced Zeolite‑Iron Oxide Nanocomposite Capacitive Biosensor for Rapid IL‑3 Sepsis Detection
- All‑Metal Terahertz Metamaterial Biosensor Achieves Ultra‑High Sensitivity for Protein Detection



