Redox‑Responsive Gelatin/Silica Nanogels Functionalized with AS1411 Aptamer for Targeted siRNA Delivery
Abstract
RNA interference (RNAi) offers unmatched specificity for gene silencing, yet siRNA stability and precise tissue delivery remain limiting factors. We engineered a dual‑responsive system by conjugating gelatin‑based nanogels (GS) with the nucleolin‑targeting AS1411 aptamer and deoxynucleotide‑substituted siRNA through a disulfide linker. This design permits transient docking of siRNA and triggers its release in a glutathione‑rich tumor microenvironment. In nucleolin‑positive A549 cells, the Apt‑GS/siRNA nanogels disassemble upon intracellular GSH exposure, efficiently delivering siRNA to the cytosol and achieving potent luciferase knock‑down. Cytotoxicity assays confirmed biocompatibility at therapeutically relevant concentrations. These redox‑responsive, tumor‑targeting nanogels represent a promising platform for siRNA‑based cancer therapy.
Introduction
RNAi harnesses sequence‑specific gene silencing, positioning it as a leading candidate for treating genetic disorders, cancers, and infectious diseases due to its precision and low toxicity. Compared with antisense oligonucleotides, small interfering RNAs (siRNAs) display enhanced nuclease resistance. Chemical modifications—such as substituting ribonucleotides with deoxynucleotides—have been shown to boost siRNA stability while retaining ~40 % of RNAi activity. Despite these advances, efficient, tissue‑specific delivery remains a formidable challenge.
Nanoparticle carriers can shield siRNA from degradation and facilitate targeted delivery. Nucleolin, overexpressed on the surface of many proliferating cancer cells, is an attractive target for tumor‑specific uptake. The G‑quadruplex DNA aptamer AS1411 (AGRO100) binds nucleolin with high affinity and has entered Phase II clinical trials as an anticancer agent. Previous studies have successfully conjugated AS1411 to various nanomaterials, achieving selective tumor cell internalization. Gelatin, a biodegradable protein, has been employed for siRNA encapsulation due to its excellent biocompatibility and gelation properties. Our group has previously developed siloxane‑crosslinked gelatin nanogels (GS NGs) with controlled size and surface charge, demonstrating efficient transfection in vitro and in vivo.
Overcoming intracellular barriers—especially endosomal escape and siRNA release—is critical. Stimuli‑responsive linkages, such as disulfide bonds that cleave in the high‑glutathione (GSH) environment of tumor cells, have attracted significant interest. Disulfide‑crosslinked carriers can trigger a burst release of therapeutic payloads specifically within malignant cells.
Here we present a hybrid nanogel system that combines GS NGs, AS1411 aptamer targeting, and a disulfide linker for redox‑triggered siRNA release. We evaluate its ability to enhance cellular uptake, achieve tumor‑specific gene silencing, and maintain low cytotoxicity.
Methods
Materials
Gelatin (Bloom 240–270, pH 4.5–5.5) was sourced from BBI Company Inc. (USA). SPDP, GPSM, and APTMS were purchased from Sigma‑Aldrich (USA). MTT, DMEM, FBS, penicillin, and streptomycin were obtained from Hyclone (USA). AS1411, 5′‑FAM‑AS1411, and the negative control TDO were synthesized by Sangon Biotech (China). Deoxynucleotide‑substituted Luc‑siRNA (sense strand: 5′‑SH‑(CH2)6‑CTTACGCTGAGTACTTCGATT‑3′; antisense strand: 3′‑TTGAAUGCGACUCAUGAAGCU‑5′; FAM‑labeled antisense) was HPLC‑purified by Gene Pharma (China). Lipofectamine 2000, the luciferase plasmid pGL3, and the luciferase assay system were from Promega (USA). A549 lung adenocarcinoma cells and NIH 3T3 fibroblasts were provided by the Biomedical Engineering Centre of Xiamen University (China). All reagents were analytical grade; glassware was thoroughly cleaned and rinsed with deionized water.
DNA sequences used:
- AS1411 aptamer: 5′‑GGTGGTGGTGGTTGTGGTGGTGGTGGTTTTTTTTTTTT‑3′
- Control TDO: 5′‑CACCGGGAGGATAGTTCGGTGGCTGTTTTTTTTTTTTT‑3′
Preparation and Characterization of Apt‑GS/siRNA Complexes
GS nanogels were synthesized via a sol‑gel process: GPSM (0.2 g) was added to 20 mL of 0.75 % gelatin in HCl (pH 3) at 60 °C for 30 min, followed by APTMS (0.08 g) and a 24‑h incubation. The resulting GS NGs were purified by centrifugation (14 000 rpm, 25 °C, 12 min) three times. Next, GS NGs (0.5 mL, 5 g L⁻¹ PBS) were reacted with SPDP (25 µL, 20 mM DMSO) for 60 min at room temperature, then incubated with thiolated AS1411 (1 mg mL⁻¹) for 12 h. The Apt‑GS nanogels were recovered by centrifugation and washed with deionized water thrice. The SPDP‑activated Apt‑GS NG suspension (80 mL, 5 mg mL⁻¹) was mixed with the siRNA sense strand overnight at 4 °C. After purification, the antisense strand was annealed by heating to 94 °C for 5 min and cooling to 47 °C for 20 min, forming Apt‑GS/siRNA complexes.
Transmission electron microscopy (TEM, Hitachi S‑4300) revealed spherical particles. Dynamic light scattering (Malvern Zetasizer) measured hydrodynamic diameters and zeta potentials. Conjugation efficiency was quantified by fluorescence of FAM‑labeled DNA and ninhydrin assay for surface amino groups (≈0.642 mmol g⁻¹).
Gel Electrophoresis Analysis
Electrophoresis was performed at 100 V for 20–30 min on 2 % agarose in TBE buffer. Images were captured with a gel documentation system. For release studies, GS/siRNA and Apt‑GS/siRNA complexes were incubated with 10 mM GSH (PBS) for 2 h prior to loading.
In Vitro Cytotoxicity
MTT assays were conducted on A549 cells (1 × 10⁴ cells / well) in 96‑well plates. After 24 h to reach 70 % confluence, cells were treated with nanogels (100–600 µg mL⁻¹) in serum‑free DMEM for 24 h, followed by 4 h incubation with MTT (5 mg mL⁻¹). DMSO was added, and absorbance was read at 490 nm. Experiments were triplicated; viability (%) was calculated relative to untreated controls.
Cell Internalization
A549 and 3T3 cells were incubated with 25 µL FAM‑labeled Apt‑GS or TDO‑GS (2 mg mL⁻¹) for 10 h in serum‑free medium at 37 °C. Cells were fixed with 4 % paraformaldehyde and imaged by confocal laser scanning microscopy (CLSM). For flow cytometry, cells were treated with FAM‑labeled nanogels (2 mg mL⁻¹) for 0.5–16 h, then washed, resuspended in ice‑cold PBS, and analyzed on an EPICS XL flow cytometer. Uptake inhibition was assessed by pre‑incubating cells with free AS1411 (0.5, 5, 50 µM) for 8 h before adding nanogels.
Gene Silencing
A549 (2 × 10⁵ cells / well) and NIH 3T3 (4 × 10⁵ cells / well) cells were transfected with Apt‑GS/siRNA complexes (80 µg mL⁻¹) for 8 h. Three treatment regimes were evaluated: (a) Apt‑GS/siRNA alone; (b) Apt‑GS/siRNA followed by liposome (A‑GS/si→Lip); (c) concurrent Apt‑GS/siRNA and liposome (A‑GS/si+Lip). After an additional 40 h, luciferase activity was measured with a luciferase assay system. Data were presented as mean ± SEM (triplicate).
Results and Discussions
Synthesis and Characterization of GS‑AS1411/siRNA Complexes
TEM images (Fig. 1) confirmed that both blank GS and Apt‑GS nanogels formed uniform spherical particles with diameters of 170–200 nm. Fluorescence imaging demonstrated successful conjugation of AS1411 and siRNA. Quantification revealed ~0.65 µmol g⁻¹ of aptamer on GS. Hydrodynamic sizes increased from 162.3 nm (GS) to 216.5 nm (Apt‑GS) and 234.9 nm (Apt‑GS/siRNA), reflecting surface functionalization. Zeta potentials shifted from +40.1 mV (GS) to +33.9 mV (Apt‑GS) and –24.9 mV (Apt‑GS/siRNA), indicating a reduction in surface charge that is expected to lower non‑specific protein binding and cytotoxicity.
siRNA Loading and Release
Gel electrophoresis showed efficient siRNA loading onto Apt‑GS/siRNA complexes, as evidenced by reduced mobility (lanes 10–11). Upon incubation with 10 mM GSH, siRNA was released within 2 h (lanes 6–7), confirming redox‑triggered disassembly. Loading capacity was estimated at 3:1 to 2:1 pmol siRNA per µg of nanogel. Physical entrapment (mixing free siRNA with GS) displayed weaker retention, underscoring the advantage of covalent attachment.
Cytotoxicity and Nucleolin Targeting
MTT assays (Fig. 4) revealed >80 % viability for GS, Apt‑GS, and Apt‑GS/siRNA across 100–600 µg mL⁻¹, demonstrating excellent biocompatibility. Slightly reduced viability with Apt‑GS was attributed to AS1411‑induced NF‑κB inhibition and organelle disruption. Fluorescence microscopy and flow cytometry (Figs. 5 & 6) confirmed selective uptake: Apt‑GS showed 96 % internalization in A549 cells versus 52 % in 3T3 fibroblasts. Competition with free AS1411 reduced uptake by 50 % (fluorescence) and 30 % (percentage of positive cells), confirming nucleolin‑mediated endocytosis.
RNA Interference
Luciferase knock‑down assays (Fig. 7a) demonstrated a 4.6‑fold reduction in A549 cells with Apt‑GS/siRNA, compared to only 6 % in NIH 3T3 cells, highlighting tumor specificity. Addition of liposomes (A‑GS/si→Lip) did not significantly enhance silencing (30.4 %) relative to Apt‑GS/siRNA alone, whereas concurrent liposome treatment (A‑GS/si+Lip) increased knock‑down to 41.6 %. The highest silencing (≈ 2.7‑fold over 3T3) was achieved at 80 µg mL⁻¹, with diminished efficiency at 120 µg mL⁻¹ likely due to increased cytotoxicity.
Conclusions
We have engineered a redox‑responsive, nucleolin‑targeted nanogel platform that protects deoxynucleotide‑modified siRNA and releases it specifically within the GSH‑rich tumor cytosol. The ~200 nm, positively charged Apt‑GS/siRNA nanogels achieve potent, tumor‑selective gene silencing (~5‑fold improvement in A549 cells) while maintaining low cytotoxicity. These findings validate the strategy of combining disulfide‑linker chemistry with aptamer targeting for efficient siRNA delivery and represent a promising step toward clinical RNAi therapeutics.
Availability of Data and Materials
The datasets supporting the conclusions of this article are included within the article.
Abbreviations
- APTMS
- 3‑Aminopropyl‑trimethoxysilane
- DMEM
- Dulbecco’s modified Eagle’s medium
- DMSO
- Dimethyl sulfoxide
- FAM
- Fluorescein amidite
- GE
- Gel electrophoresis
- GPSM
- 3‑Glycidoxypropyl‑trimethoxysilane
- GS NGs
- Siloxane‑crosslinked gelatin nanogels
- MTT
- 3-(4,5‑Dimethylthiazol‑2‑yl)-2,5‑diphenyltetrazolium bromide
- RNAi
- RNA interference
- siRNAs
- Small interfering RNAs
- SPDP
- N‑succinimidyl 3-(2‑pyridyldithio)‑propionate
- TDO
- Negative DNA oligonucleotide
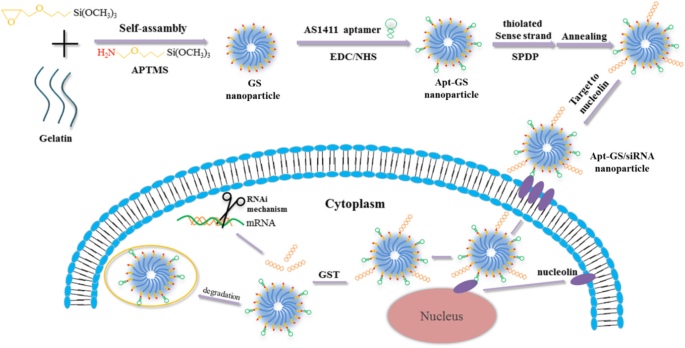
A redox‑responsive and tumor‑targeting Apt‑GS nanogels were developed for siRNA delivery with disulfide conjugation and aptamer AS1411.
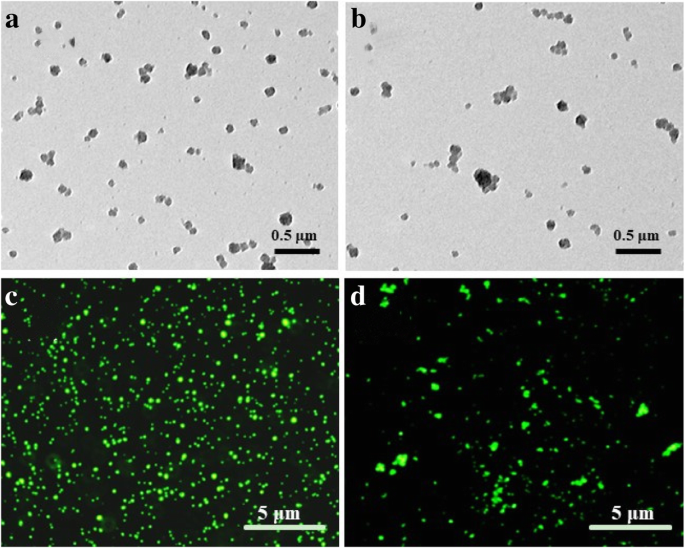
Images of Apt‑GS (a, c) and Apt‑GS/siRNA nanogels (b, d). a, b TEM images. Scale bars: 0.5 µm. c, d Fluorescent images. Scale bars: 5 µm.
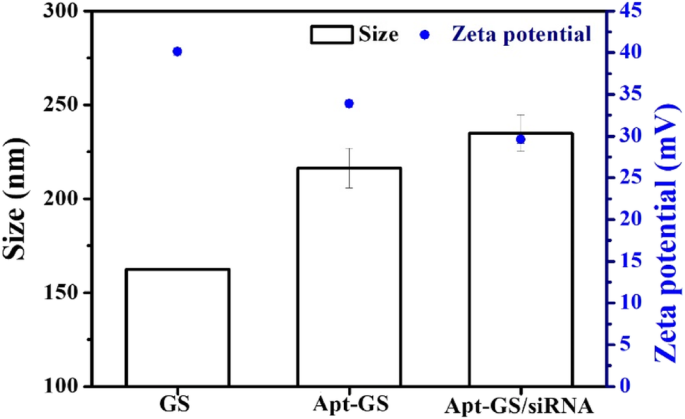
Hydrodynamic size and zeta potentials of GS, Apt‑GS, and Apt‑GS/siRNA nanogels.
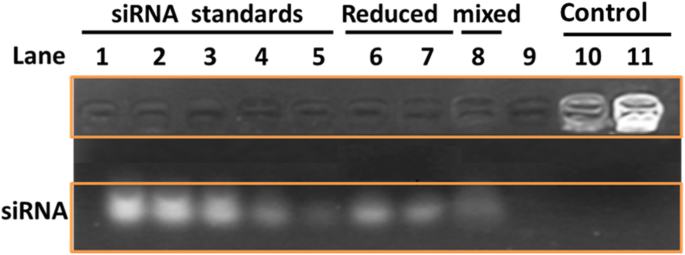

AGE analysis of siRNA reversible binding to Apt‑GS nanogels. Lanes 1–5: free siRNA (50–10 pmol). Lanes 6–7: GS/siRNA and Apt‑GS/siRNA with 10 mM GSH (2 h). Lane 8: free siRNA mixed with Apt‑GS NGs. Lanes 10–11: GS/siRNA and Apt‑GS/siRNA without GSH.
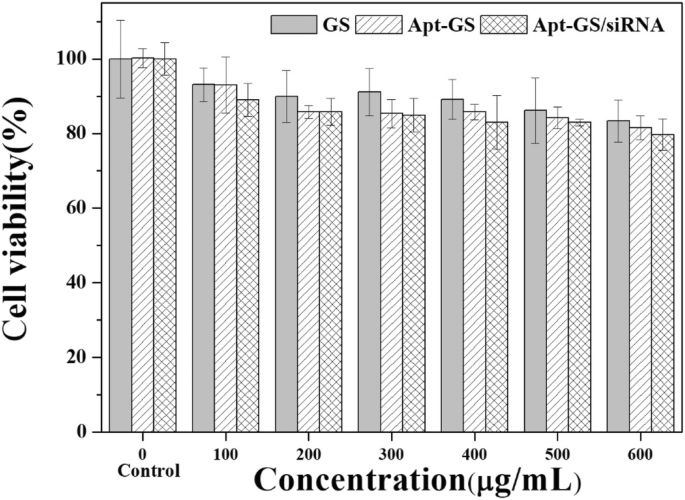
Cell viability of A549 cells with GS, Apt‑GS, and Apt‑GS/siRNA nanogels at various concentrations (MTT assay).
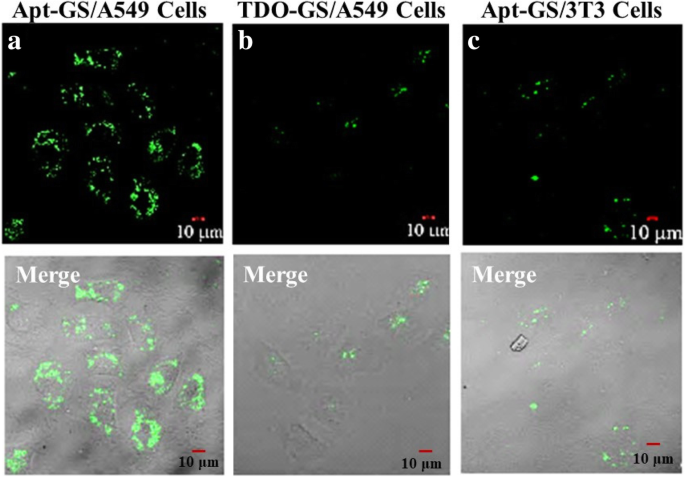
a–c Cell uptake of FAM‑siRNA delivered by Apt‑GS and TDO‑GS nanogels. Images from CLSM. Green signal: FAM. Scale bar: 10 µm.
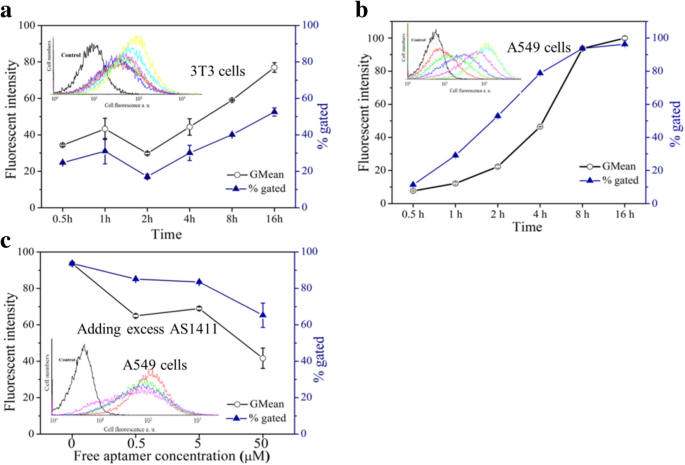
Flow cytometry analysis of cell uptake of FAM‑labeled Apt‑GS and TDO‑GS nanogels. Insert: FACS profiles. a, b curves: incubation time (0.5–16 h). c curve: AS1411 concentration (0–16 µM).
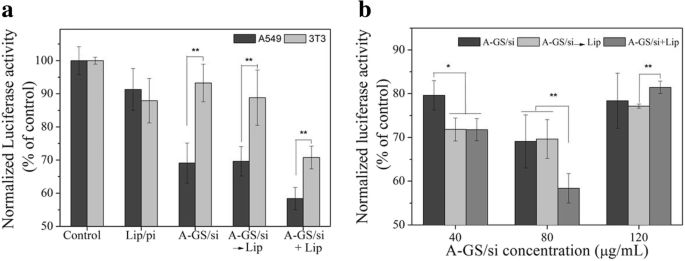
In vitro silencing effect of anti‑luciferase siRNA formulated in Apt‑GS nanogels with different treatments (a) and concentration of Apt‑GS/siRNA complexes (b) on gene silencing.
Nanomaterials
- Nanofiber & Filament-Based Nanocarriers: Advancing Precision Drug Delivery
- Cell‑Based Drug Delivery Systems for Advanced Cancer Therapy
- 131I‑Labeled PLGA‑Lipid Nanoparticles for Targeted Paclitaxel Delivery and Imaging in Melanoma Therapy
- Innovative Nucleoside–Lipid Nanocarriers Enhance Sorafenib Delivery and Reduce Toxicity
- Reimagining Nanoparticle Therapies Beyond Cancer
- Targeted Cancer Delivery: Decorating Nanovesicles with pHLIP for Acidic Tumor Microenvironment Recognition
- Targeted Delivery of Doxorubicin Using Biocompatible Chitosan Nanobubbles and Ultrasound
- Efficient Colon Cancer Gene Therapy Using Cationic DOTAP–PEG‑PCL Micelles for siRNA Delivery
- Biotin‑Functionalized Redox‑Sensitive Chitosan Nanoparticles Coated with Phycocyanin for Targeted Curcumin Delivery
- Targeted siRNA Delivery to Ovarian Cancer Using Folate‑Functionalized Graphene Oxide/PEI Nanocomplexes



