Pharmacophore‑Guided Virtual Screening Enhances Drug Repurposing for COVID‑19
Drug repurposing—reassigning approved or investigational drugs to new therapeutic uses—has long been a cornerstone of rapid therapeutic discovery.1 Its most celebrated success, the transformation of a hypertension and angina medication into Pfizer’s “little blue pill,” Viagra, showcases its potential impact. In the wake of COVID‑19, this strategy has again captured global attention.
Three‑dimensional protein modeling offers several in‑silico tools for drug repurposing. A prior study in this series demonstrated how molecular docking can identify potential inhibitors of the SARS‑CoV‑2 main protease (Mpro). Here, we ask whether pharmacophore modeling—an abstraction of the essential ligand–protein interaction features—can deliver comparable insights and add further confidence in selecting candidate molecules.
Pharmacophore Modeling as an Alternative Approach
Pharmacophore models distill the key non‑bonding interactions that enable a ligand to bind a protein target, offering a complementary perspective to traditional simulation methods. We began with a set of SARS‑CoV‑2 Mpro crystal structures obtained from the Diamond Light Source.2 After aligning the structures, we employed BIOVIA Discovery Studio’s “Interaction Pharmacophore Generation” protocol to extract pharmacophore features from each protein‑ligand complex. Feature counts ranged from two (for 5R80 and 5R7Y) to nine (for 6LU7). By merging and simplifying overlapping features, we arrived at a 14‑feature consensus pharmacophore that encapsulates the critical intermolecular contacts of the protease with potential binders.
To refine this model, we performed in‑silico alanine‑scanning mutagenesis on the active‑site residues across all complexes. Eight residues—HIS41, MET49, ASN142, HIS163, MET165, PRO168, GLN189, and GLN192—emerged as hotspots in at least one complex, with three residues (HIS41, MET49, MET165) identified as hotspots in four or more complexes. Six of the 14 pharmacophore features directly mapped onto interactions with these hotspot residues. A second dataset of non‑covalent Mpro ligand complexes revealed alternative binding modes, prompting us to separate the six key features into two groups for subsequent virtual screening.
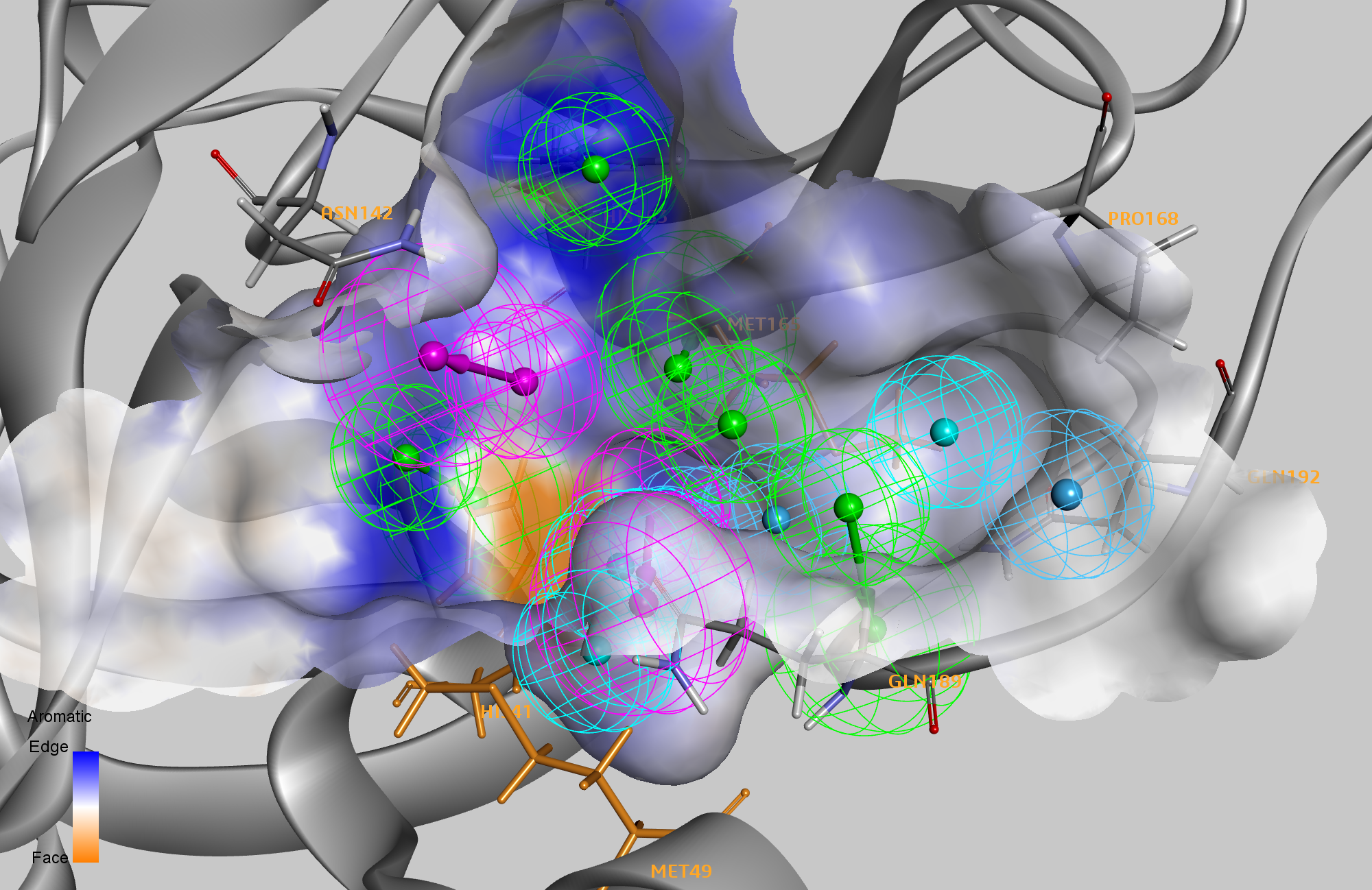
Using this refined pharmacophore, we screened a library of 2,650 FDA‑approved drugs, each pre‑generated with multiple ligand conformations to accelerate searching. Hits were defined by matching at least one feature from each of the two essential groups and a minimum of two additional features from the remaining eight. This stringent criterion ensured diverse exploration of binding modes while preserving interactions with critical residues identified via alanine scanning. The screening algorithm explored over 12,000 pharmacophore combinations. For each hit, we performed energy minimization within the protein context and calculated binding energies using CHARMm with a Generalized Born/Volume (GBMV) implicit solvent model—effectively a docking calculation constrained by the pharmacophore.
We focused on poses that engaged four key residues—HIS41, MET49, CYS145, and MET165—because these comprise the catalytic dyad (HIS41/CYS145) and are common to multiple hotspot residues. Sorting by calculated binding free energy yielded the top ten candidates. Ritonavir, already highlighted in our earlier docking study, ranked seventh in both analyses and is currently in several COVID‑19 clinical trials.3
Video 1: Best‑scoring pose of Ritonavir interacting with HIS41, MET49, SER144, CYS145, MET165, GLU166, and GLN189.
The leading ligand from this pharmacophore‑driven approach was Montelukast, a cysteinyl leukotriene receptor antagonist. Recent literature suggests its potential to mitigate COVID‑19 progression.4
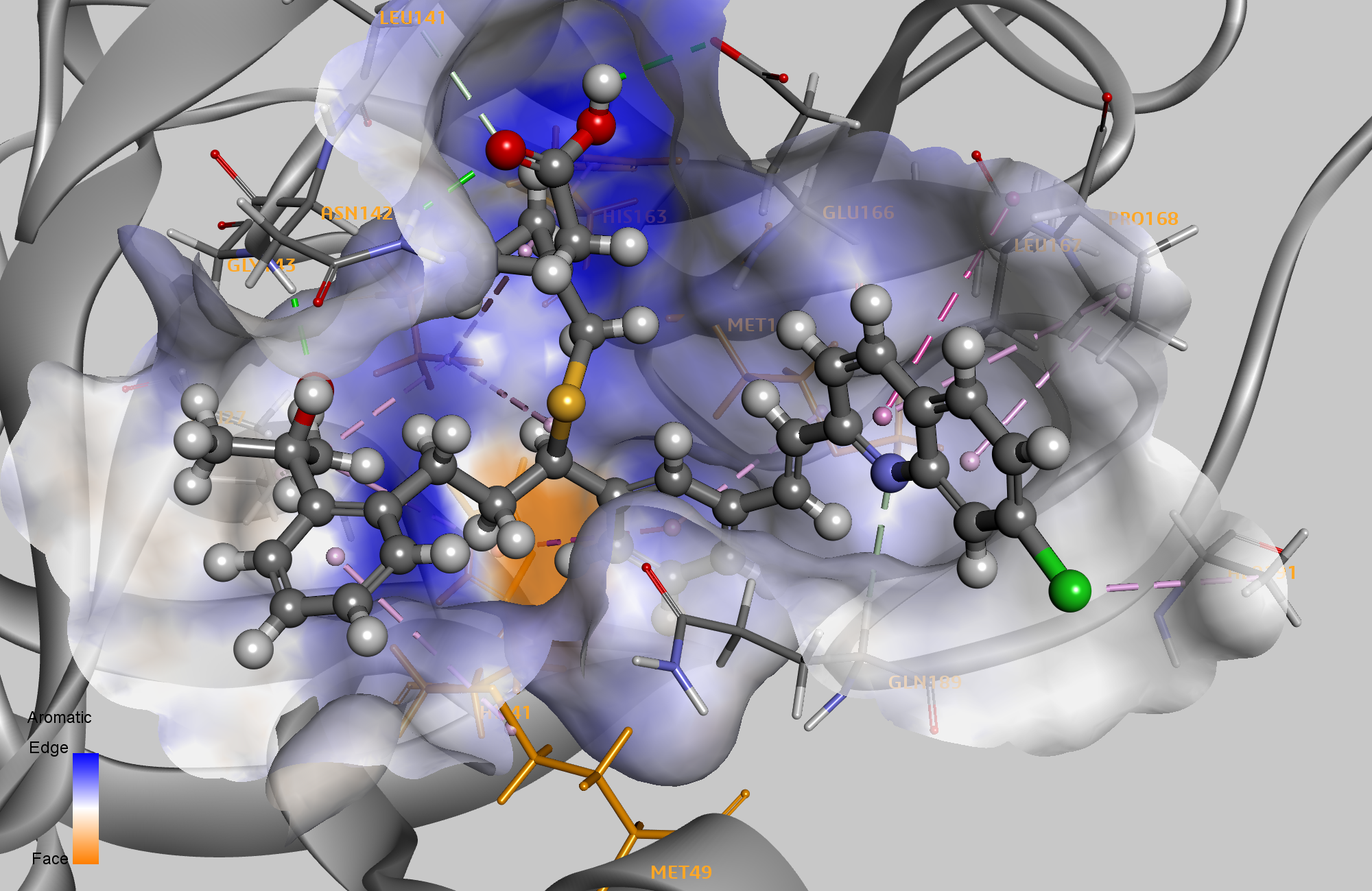
Other top‑scoring compounds included Telmisartan, Moexipril, and hydroxychloroquine—each recently scrutinized for repurposing potential, with hydroxychloroquine generating the most controversy.5 Notably, the pharmacophore‑derived hit list showcased a broader range of target classes compared to the docking‑only list, which was dominated by HCV or HIV protease inhibitors.
Our results demonstrate that integrating in‑silico alanine‑scanning mutagenesis into pharmacophore modeling can refine virtual screening, yielding a prioritized set of diverse drug candidates. While the overlap with docking‑derived leads was modest, the shared identification of Ritonavir underscores the value of consensus approaches. Combining the strongest candidates from both methods may guide more focused experimental validation.
In summary, pharmacophore‑guided virtual screening serves as a powerful, complementary strategy to docking, enhancing confidence in candidate selection and expanding the chemical space explored for COVID‑19 therapeutics.
Biologics
- Virtual Inventories & 3D Printing: Securing the Future of Distributed Manufacturing
- Nanofiber & Filament-Based Nanocarriers: Advancing Precision Drug Delivery
- Repurposing FDA‑Approved Drugs to Inhibit SARS‑CoV‑2 Main Protease: A Virtual Screening Analysis
- Cell‑Based Drug Delivery Systems for Advanced Cancer Therapy
- Chitosan‑Capped, Enzyme‑Responsive Hollow Mesoporous Silica Nanoplatforms for Targeted Colon Drug Delivery
- Liposomal Nanomedicine for Targeted Cancer Drug Delivery: Enhancing Efficacy and Safety
- CompositesWorld Partners with ITHEC for Expanded 3-Day Virtual Thermoplastic Composites Conference
- Universal Robots Unveils the World’s Largest Virtual Conference on Collaborative Robots
- 5 Expert Tips to Ace Your Virtual Job Interview – Infographic
- Tiny 3D-Printed Micro-Robots Pave the Way for Precision Drug Delivery



