Unraveling the Origins of SARS‑CoV‑2: A Genomic Perspective
The COVID‑19 pandemic remains a formidable threat to global public health and has disrupted economies worldwide. SARS‑CoV‑2, the virus responsible for the outbreak, is not a typical influenza strain; it targets both upper and lower respiratory tracts, impairing essential breathing processes and causing significant mortality. As of April 6, 2020, Worldometer reported 1,337,166 confirmed cases and 74,176 deaths globally.
Analyzing the SARS‑CoV‑2 genome provides critical insights into the virus’s origin, informs diagnostic development, and guides therapeutic innovation to reduce loss of life.
Understanding the SARS‑CoV‑2 Genome
SARS‑CoV‑2 is a single‑stranded RNA virus with a genome of nearly 30 kilobases and 12 putative open reading frames. Chinese researchers first sequenced the genome in December 2019, and multiple teams have since released complete sequences, now publicly available in GenBank and the Coronavirus Database.
Origin of the SARS‑CoV‑2 Virus
Conspiracy theories often arise during outbreaks, fostering prejudice against nations, cultures, and communities. A rigorous, evidence‑based approach is essential to counter misinformation. Genome analyses unequivocally show that SARS‑CoV‑2 evolved naturally and is not a laboratory construct.1,2 More than 100 strains sampled worldwide are over 99.5% identical at the nucleotide level, indicating minimal regional mutation and a high transmissibility.
Previous coronaviruses—SARS‑CoV (2002, China) and MERS‑CoV (2012, Saudi Arabia)—were traced to bats. Sequencing bat coronavirus RaTG13 revealed a 96.2% identity to SARS‑CoV‑2, confirming a zoonotic origin. Additionally, a 91% genomic similarity to Malayan pangolin coronavirus suggests pangolins may have served as an intermediate host. Pangolins, prized in Asia for traditional medicine and as delicacies, are the most trafficked mammal worldwide.
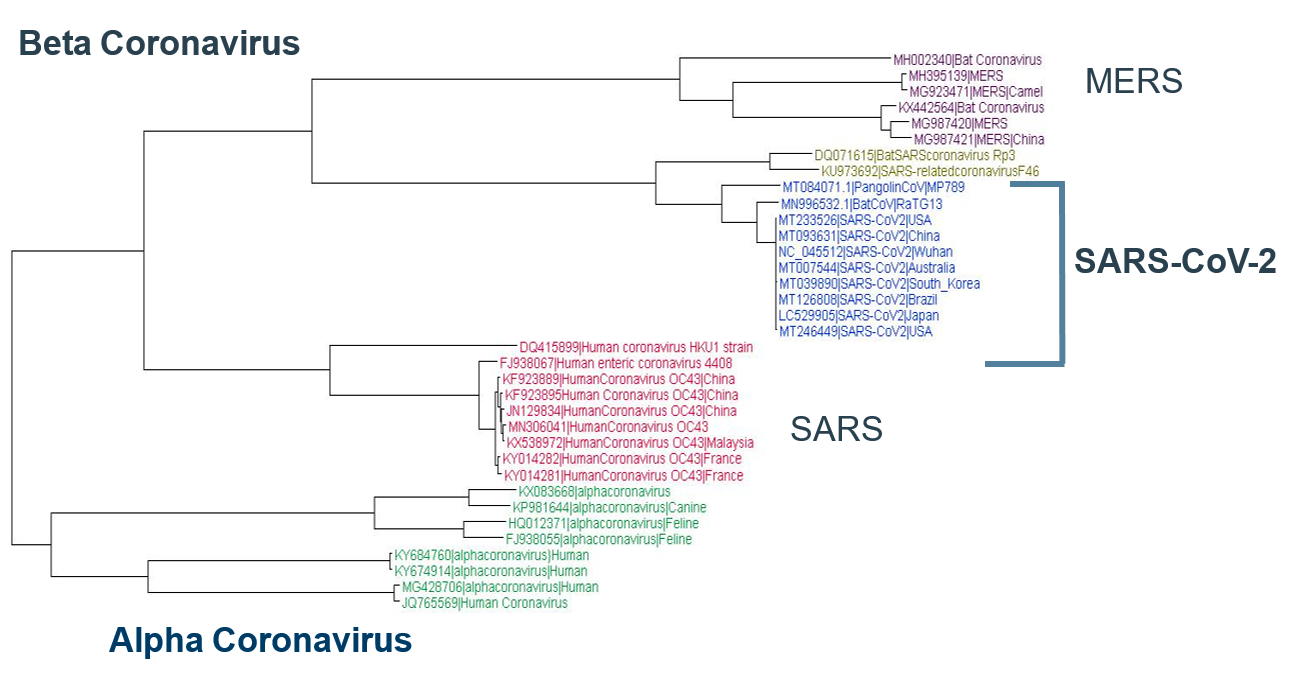
SARS‑CoV‑2 differs from other coronaviruses by possessing 88% or less sequence identity. Phylogenetic analyses place SARS‑CoV‑2, RaTG13, and pangolin CoV within a novel clade of beta‑coronaviruses, as illustrated below.
How Does the Coronavirus Enter the Host?
The spike (S) protein is a multifunctional molecular machine composed of two subunits, S1 and S2. S1 binds the host receptor, while S2 mediates membrane fusion. The receptor‑binding domain (RBD), 193 amino acids long, attaches to the human ACE2 receptor—expressed on lung, arterial, cardiac, renal, and intestinal cells. ACE2 normally regulates blood pressure by converting angiotensin II to angiotensin 1‑7 but also serves as a viral entry point.
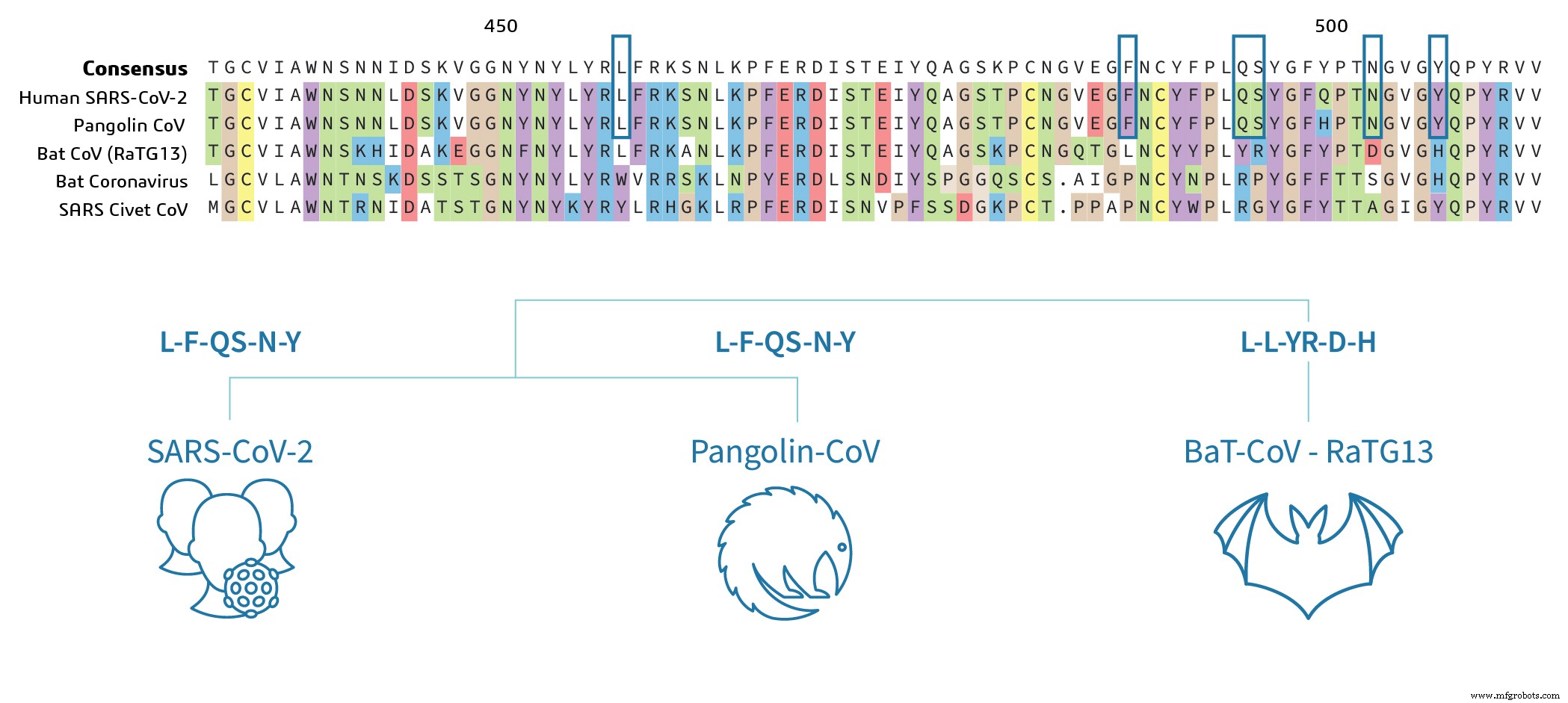
Pangolin CoV and SARS‑CoV‑2 share highly conserved RBD sequences, including identical key binding residues (LFQSNY). In contrast, the bat SARS‑CoV‑2 RBD differs at 17 residues, including five critical binding sites, suggesting limited ACE2 interaction without experimental confirmation.
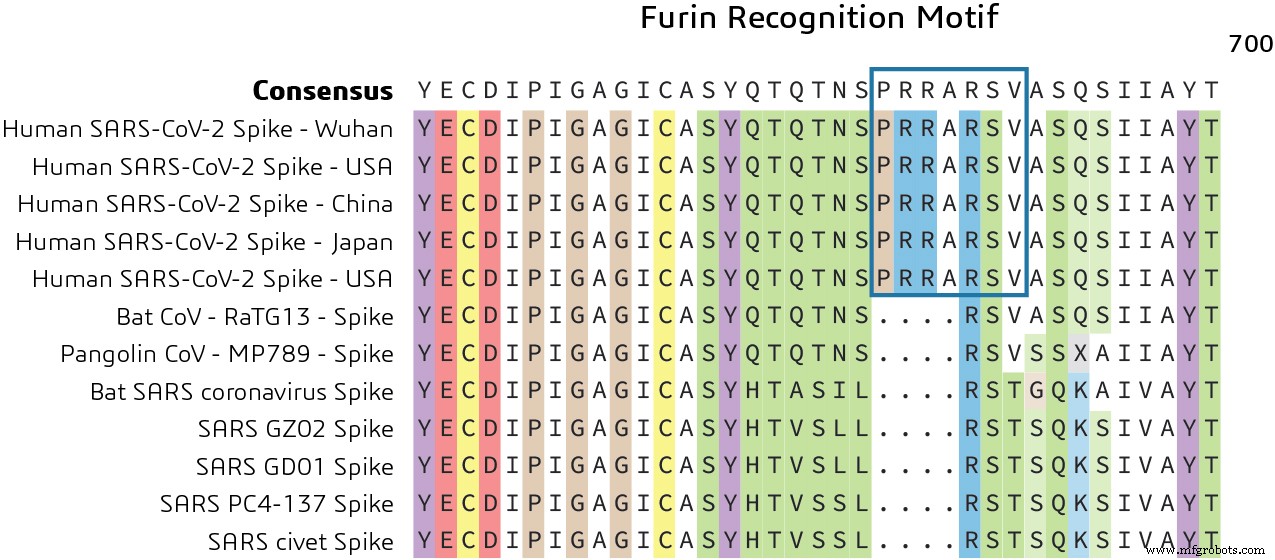
The spike protein also contains a furin cleavage site at the S1/S2 boundary, denoted PRRARSV, which is recognized by the host FURIN protease. Acquisition of this motif may represent a ‘gain‑of‑function’ event that enabled the virus to efficiently infect human cells. Targeting this cleavage site could offer a therapeutic strategy to block viral entry.
Overall, the spike protein’s optimized RBD, furin cleavage motif, and strong ACE2 affinity suggest natural selection and recombination events between bat and pangolin coronaviruses. These findings support a zoonotic origin generated through natural recombination rather than laboratory manipulation.
Immediate Donor of SARS‑CoV‑2 to Humans
The SARS‑CoV‑2 genome exhibits a mosaic of bat RaTG13 and pangolin CoV sequences, consistent with recombination. Identifying the precise intermediate host—whether another pangolin species or a different wildlife species present in the Wuhan seafood market—remains an open question. Clarifying the origin is crucial for preventing future spillover events and global pandemics.
For further information, contact us at https://www.3dsbiovia.com/about/contact/.
Biologics
- How COVID‑19 is Reshaping Manufacturing: Production, Demand, and Supply Chain Impacts
- Decoding SARS‑CoV‑2 Genomes: DNA & Antibody Tests for COVID‑19
- Leveraging Supply Chain Innovation to Combat COVID‑19
- How COVID-19 Is Disrupting Global Supply Chains
- RapidPlex Sensor: 5 Essential Questions About SARS-CoV-2 At‑Home Testing
- Discover the Crippa Tube Bender: Italy’s Leading Tube Bending Innovation
- Understanding Aluminum Alloy Designations: A Practical Guide
- The Evolution of Sawing: From Brass Reinforcements to Steel Innovations
- How COVID‑19 Reshaped Industrial Automation: Trends, Challenges, and Opportunities
- Mastering Duty Cycle Ratings of Piston Compressors



