Ultra‑Sensitive Electrochemical DNA Biosensor Using Acrylic‑Gold Nanocomposite for Rapid Arowana Fish Gender Determination
Abstract
This study introduces an ultra‑sensitive electrochemical DNA biosensor that identifies the gender of the Asian arowana (Scleropages formosus) with an atto‑M detection limit. The sensor employs acrylic microspheres functionalized with succinimide groups and colloidal gold nanoparticles (AcMP‑AuNP) immobilized on a carbon screen‑printed electrode. An aminated DNA probe is covalently anchored to the AcMPs, and target arowana DNA hybridization is quantified by differential pulse voltammetry (DPV) using anthraquinone‑sulfonic acid as a redox label. Under optimized conditions the sensor reaches a detection limit of 1.0 × 10⁻¹⁸ M and a linear range of 1.0 × 10⁻¹⁸ – 1.0 × 10⁻⁸ M (R² = 0.99) within one hour, making it a lightweight, cost‑effective tool for on‑site fish‑culture diagnostics.
Background
The Asian arowana (Scleropages formosus) is a freshwater species prized for its ornamental value and cultural significance across Southeast Asia and parts of Oceania. Owing to its high market demand, wild populations have been heavily exploited, leading to its classification as endangered by CITES and the IUCN Red List. Accurate gender determination at the juvenile stage is critical for breeding programs and market regulation, yet traditional visual methods are unreliable and time‑consuming, and conventional PCR approaches are laborious and require sophisticated laboratory infrastructure.
Recent advances in nanomaterial‑based biosensors offer a promising alternative. By integrating acrylic polymer microspheres with gold nanoparticles, we can create a high‑surface‑area, conductive platform that enhances probe loading and electron transfer, thereby improving sensitivity and reducing assay time.
Methods
Apparatus and Electrodes
DPV measurements were carried out on an Autolab PGSTAT 12 potentiostat using a carbon screen‑printed electrode (SPE) modified with AcMPs and AuNPs. A platinum auxiliary electrode and an Ag/AgCl reference electrode (3.0 M KCl) were employed. Solutions were sonicated in an Elma S30H bath to ensure homogeneity.
Chemicals
Key reagents included 2‑2‑Dimethoxy‑2‑phenylacetophenone (DMPP), 1,6‑Hexanediol diacrylate (HDDA), n‑butyl acrylate (nBA), Au(III) chloride trihydrate, and NAS. Colloidal gold nanoparticles were synthesized per Grabar et al. (1995). DNA probes were prepared in 0.05 M K‑phosphate buffer (pH 7.0) and hybridized in 0.05 M Na‑phosphate buffer (pH 7.0) containing 1.0 mM AQMS and 2.0 M NaCl.
Synthesis of Acrylic Microsphere
AcMPs were produced via one‑step UV‑initiated photopolymerization of a mixture containing HDDA, nBA, DMPP, and NAS in an aqueous emulsion. After 10 min sonication and 600 s UV curing under N₂, the microspheres were collected by centrifugation, washed thrice with K‑phosphate buffer, and air‑dried.
Fabrication of DNA Biosensor Using Acrylic Microspheres
The SPE surface was first drop‑coated with a 3 mg/mL AcMP dispersion, then with 5 mg/mL colloidal AuNPs. The modified electrode was incubated in a 5 µM aminated arowana DNA probe solution for 6 h, followed by thorough rinsing. Hybridization with target cDNA (5 µM) in the presence of 1 mM AQMS and 2 M NaCl proceeded for 1 h before DPV analysis in 0.05 M K‑phosphate buffer at room temperature.
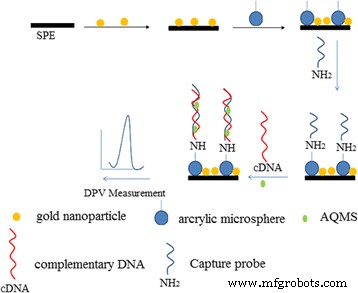
The fabrication procedure of the electrochemical arowana DNA biosensor based on AcMP‑AuNP‑modified electrode.
Optimization of Electrochemical Arowana DNA Biosensor
Probe immobilization was optimized by varying incubation times (1–18 h) and hybridization times (10–100 min). Temperature, pH, ionic strength, and cation valency were systematically examined to maximize DPV response. The optimal conditions were: 6 h probe incubation, 30 min hybridization at 25 °C, pH 7.0, 0.05 M Na‑phosphate buffer, 2 M NaCl, and 1 mM AQMS.
DNA Extraction and Arowana DNA Analysis
Fifteen arowana tissue samples were extracted using the QIAquick PCR Purification kit. Genomic DNA was amplified by PCR and verified on 1.5 % agarose gels. The same extracts were tested with the biosensor; DPV currents above baseline indicated male DNA, while currents at baseline corresponded to female DNA. Statistical significance was assessed using a t‑test (95 % confidence).
Results and Discussion
SEM images confirmed uniform, spherical AcMPs (Fig. 2) with a narrow size distribution (Fig. 3). Electrochemical characterization of the modified SPE demonstrated a diffusion‑controlled, reversible redox process with a linear relationship between peak current and the square root of scan rate (R² = 0.996; Fig. 4). The DPV response (Fig. 6) showed a pronounced current increase only after hybridization with complementary DNA, confirming probe specificity.

SEM image of acrylic polymer microspheres.
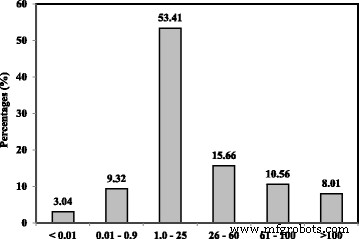
Size distribution of acrylic microspheres prepared from photopolymerization.
Optimization studies revealed that probe immobilization plateaued after 6 h, while hybridization reached equilibrium within 30 min at room temperature (Fig. 7). Temperature above 25 °C accelerated hybridization but risked denaturation, so 25 °C was chosen as optimal. pH dependence showed maximal DPV signal at neutral pH 7.0; lower or higher pH reduced hybridization efficiency (Fig. 9). Among cations tested, Na⁺ provided the best enhancement, likely due to minimal precipitation and favorable electrostatic shielding (Fig. 10). Ionic strength analysis identified 2 M NaCl and 0.05 M Na‑phosphate buffer as the optimal hybridization buffer (Fig. 11).
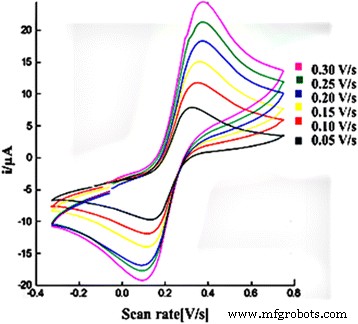
Cyclic voltammograms of 1.0 mM K₃Fe(CN)₆ in 0.05 M Na‑phosphate buffer (pH 7.0) at various scan rates for a modified SPE containing AcMP‑AuNP material.
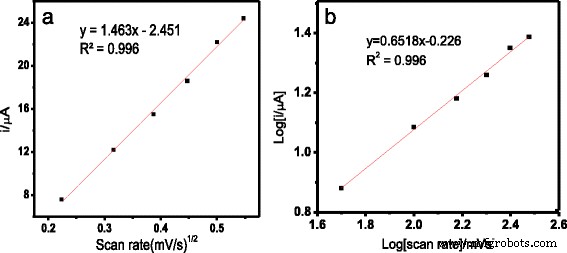
Plot of oxidation peak currents versus the square root of scan rate (a) and versus log of scan rate (b).
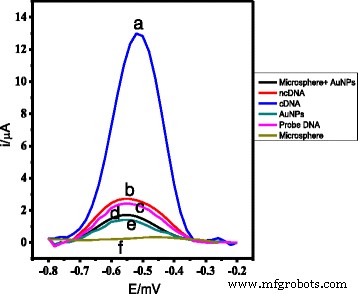
DPV signal of AcMP‑AuNP‑based DNA electrode before and after hybridization with complementary (a) and non‑complementary (b) DNA; comparison with electrodes modified only with AcMPs (f), only AuNPs (e), and AcMP‑AuNP composite (d).
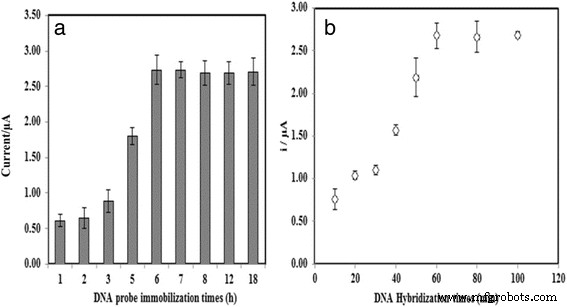
Effects of DNA probe immobilization time (a) and hybridization time (b) on the biosensor response.
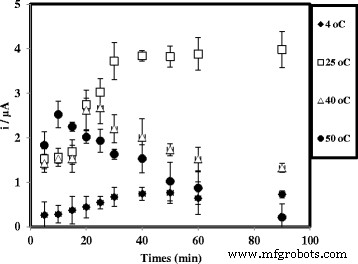
Effect of temperature on hybridization time.
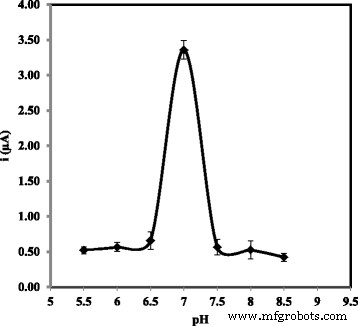
DPV response versus pH (5.5–8.0).
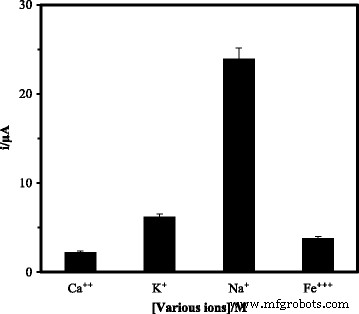
Effect of cations (Ca²⁺, Na⁺, K⁺, Fe³⁺) on DPV response.
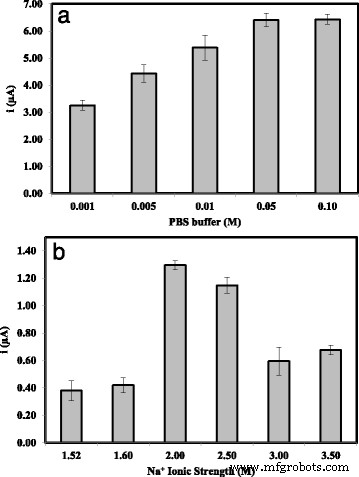
Influence of Na‑phosphate buffer concentration (a) and ionic strength (b) on biosensor response.
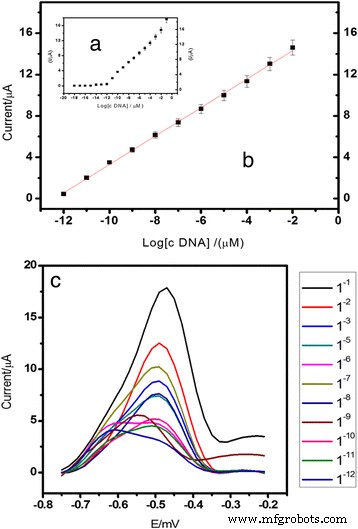
Calibration curve showing a linear response from 1.0 × 10⁻¹⁸ M to 1.0 × 10⁻⁸ M (R² = 0.99) and representative DPV spectrum.
Determination of Arowana Fish Gender with DNA Biosensor
When applied to field samples, the biosensor's gender assignments matched PCR results (Table 2), confirming its reliability for rapid, on‑site use.
Conclusions
The developed electrochemical DNA biosensor offers exceptional sensitivity, a broad dynamic range, and a low detection limit for arowana DNA. Its rapid, atto‑M level detection capability, combined with a lightweight, disposable format, makes it ideal for integration into point‑of‑use devices that can support fish‑culture operations by enabling early gender and color identification.
Nanomaterials
- IBM and Warwick Scientists Capture First High‑Resolution Image of Triangulene, a Highly Reactive Triangular Molecule
- A Flexible Graphene Biosensor for Rapid, Highly Sensitive Detection of Ovarian Cancer Cells
- N,N‑Dimethylformamide Modulates MXene Quantum Dot Fluorescence for Ultra‑Sensitive Fe³⁺ Detection
- Ultra‑Sensitive Electrochemical DNA Biosensor for Vibrio cholerae Detection Using Polystyrene‑Acrylic Acid Nanoparticles and Gold Nanoparticles
- Wireless Magnetoelastic Nanobiosensor Achieves Ultra‑Sensitive Atrazine Detection
- Enhanced Formaldehyde Detection at ppb Levels Using Reduced Graphene Oxide–Coated Silicon Nanowire Sensors
- Magnetic Graphene Field‑Effect Transistor Biosensor for Ultra‑Sensitive Single‑Strand DNA Detection
- CNT@Li₂FeSiO₄@C Core‑Shell Heterostructure: A Robust Cathode for High‑Performance Lithium‑Ion Batteries
- Ultra‑Sensitive Biosensing Using 2‑D Hole‑Array Grating‑Coupled Bloch Surface Waves
- High-Precision Flexible Temperature Sensors for Human Body Monitoring: A Comprehensive Review



